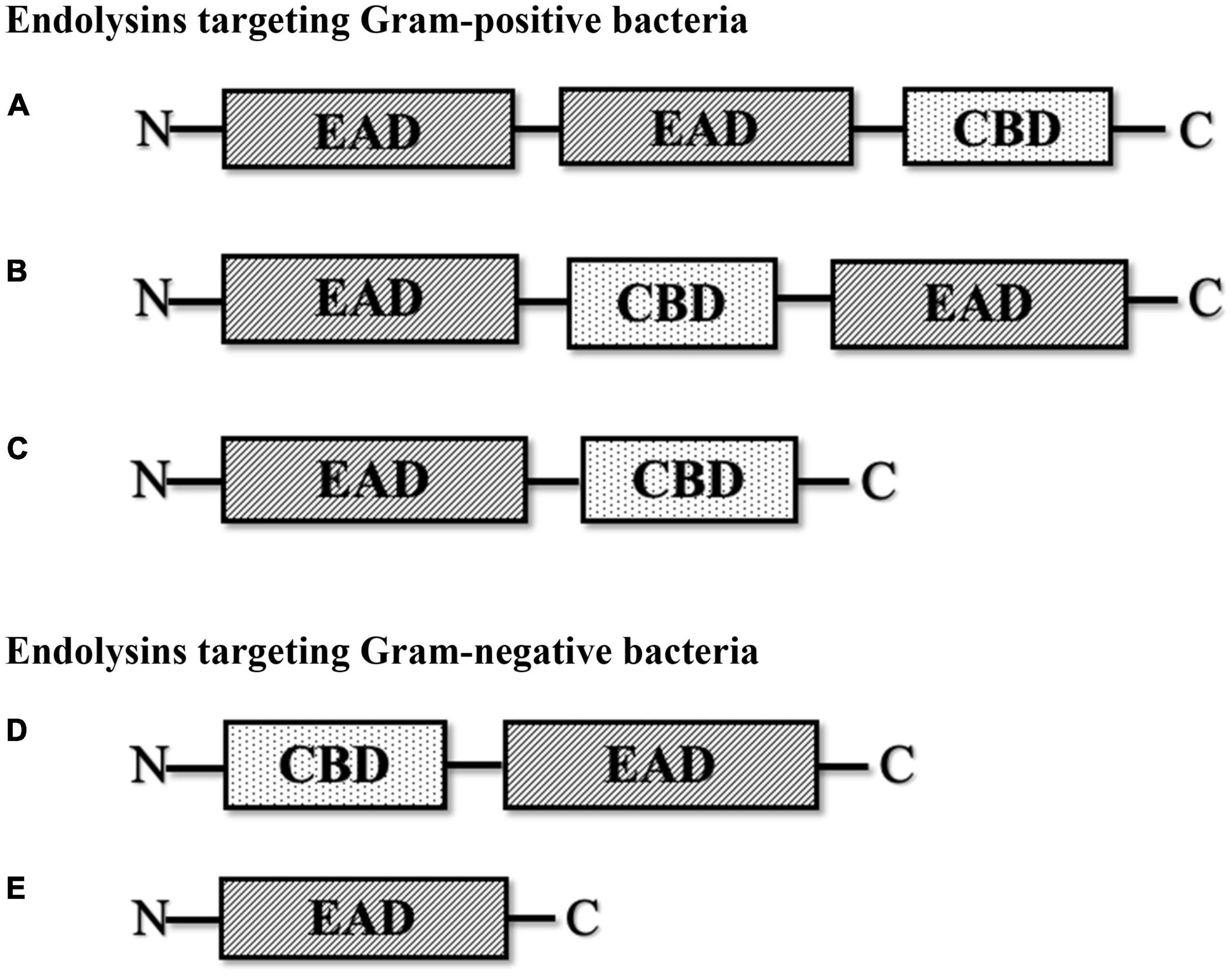
Modular configuration models of common phage endolysins. (a) Model with... | Download Scientific Diagram

Precise Photothermal Treatment of Methicillin-Resistant S. aureus Infection via Phage Lysin-Cell Binding Domain-Modified Gold Nanosheets | ACS Applied Materials & Interfaces

Bacteriophages for detection and control of foodborne bacterial pathogens—The case of Bacillus cereus and their phages - Abraha - Journal of Food Safety - Wiley Online Library
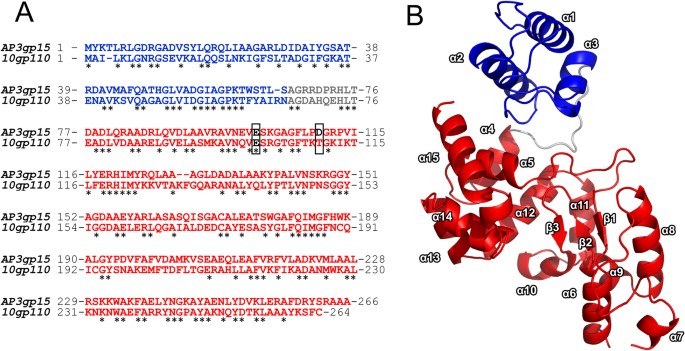
Modular endolysin of Burkholderia AP3 phage has the largest lysozyme-like catalytic subunit discovered to date and no catalytic aspartate residue | Scientific Reports

Rapid Multiplex Detection and Differentiation of Listeria Cells by Use of Fluorescent Phage Endolysin Cell Wall Binding Domains | Applied and Environmental Microbiology
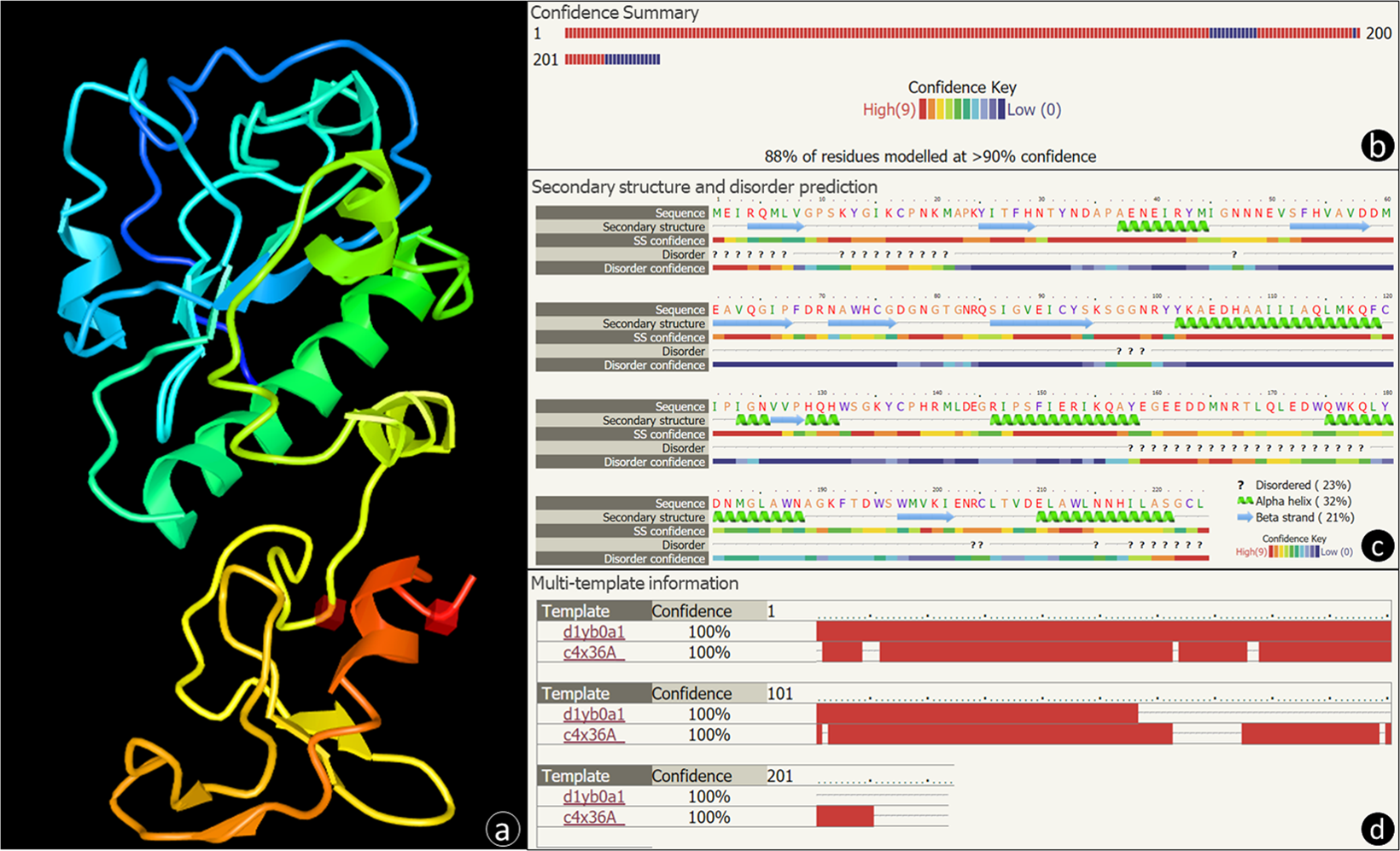
Identification of the first endolysin Cell Binding Domain (CBD) targeting Paenibacillus larvae | Scientific Reports

Frontiers | Development of a Novel Chimeric Endolysin, Lys109 With Enhanced Lytic Activity Against Staphylococcus aureus

Viruses | Free Full-Text | Reporter Phage-Based Detection of Bacterial Pathogens: Design Guidelines and Recent Developments
GitHub - ku-cbd/PhageBoost: Rapid discovery of novel prophages using biological feature engineering and machine learning